|
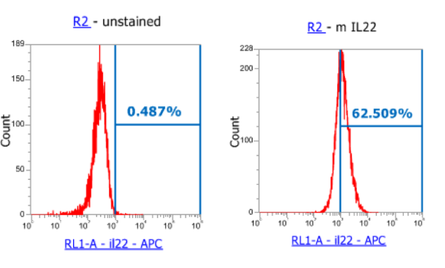
Once the 4T1 cells were put into the flow cytometer, the cells were divided into the positive and negative cultures and represents which cells are positive and negative with the presence of IL-22 in the cell cultures. In Figure 1, it is shown that in an unstained 4T1 cell culture, there is approximately 0.487% auto fluoresce in the IL-22 channel. The unstained cells were used as a control for the experiment, that way a differential could be made between stained (with tdTomato) and unstained cells that express IL-22. After the unstained cells were seen to be the control, the stained 4T1 cells for IL-22 were run through the flow cytometer. With this, which is displayed in Figure 1, the 4T1 cells that were stained to show expression of IL-22 in the culture, showcased that approximently 62.51% of the 4T1 cells fluoresced, resulting in 38.49% of the cancer cells not expressing IL-22. From the data, it was concluded that over half of the cell population expressed IL-22 and that the stain did work, resulting that IL-22 was indeed knocked out of the cell line using CRISPR.
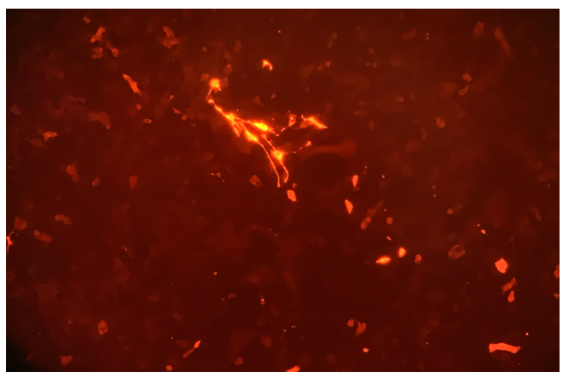
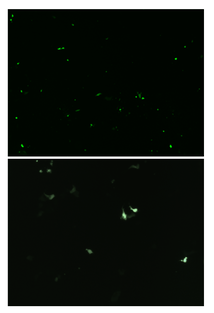
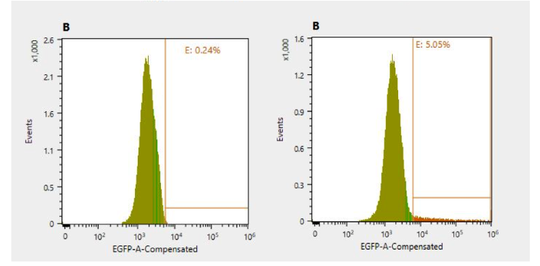
Once basal IL-22 levels were established in the 4T1s the CRISPR vector was transfected into the cells and the GFP (Green Fluorescent Protein) gene was studied in the fluorescence using a confocal microscope. The confocal microscope was used to verify the presence of the plasmid, as any cell expressing GFP would have taken up the plasmid. Seen in Figure 2, the spots of green are cells that express the modification of the cells from CRISPR. To resolve the finding of the percent of cells expressing GFP, the cell sorter was used once again. Through the cell sorter, a control culture of 4T1 cells presented that 0.24% of the cells autofluorescence in the GFP channel. A second culture containing green 4T1 cells was placed in the cell sorter and identified that 5.05% of the cells expressed GFP in the cell line (Figure 3). 5.05% of green cells that expressed GFP in the cell line represent how many cells adopted the modified plasmid, being the GFP. The 4T1 were not only stained green to check for GFP expression, but were also stained with tdTomato. Using tdTomato is will be very beneficial in the future studies for tracking metastasis to distant organs, as the cells could be removed again and verified using the dye associated with tdTomato. tdTomato also would expedite further research with the dye and cells associated with it, as results of the tumor can be collected once the tumor has become metastatic. seeing the dye on cells using a confocal microscope depicts that the stain visibly worked, and would be used in future applications and research since it is easily identifiable (Figure 4). Figure 4: tdTomato through a confocal microscope, used to depict what the stain will display on cells
|
Figure 1: unstained 4T1 cancer cells, as a control, compared to stained 4T1 cells for IL-22
Figure 3: unstained 4T1 cells as a control, compared to green stained 4T2 cells, with 5.05% expressing GFP in the cell line
|